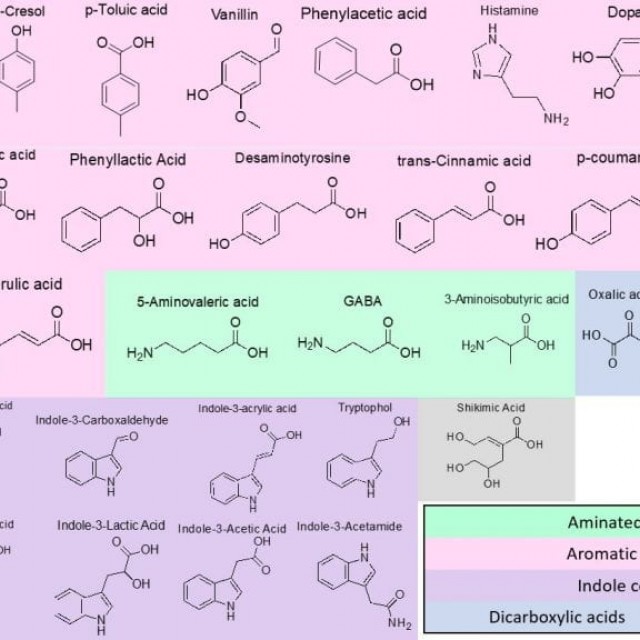
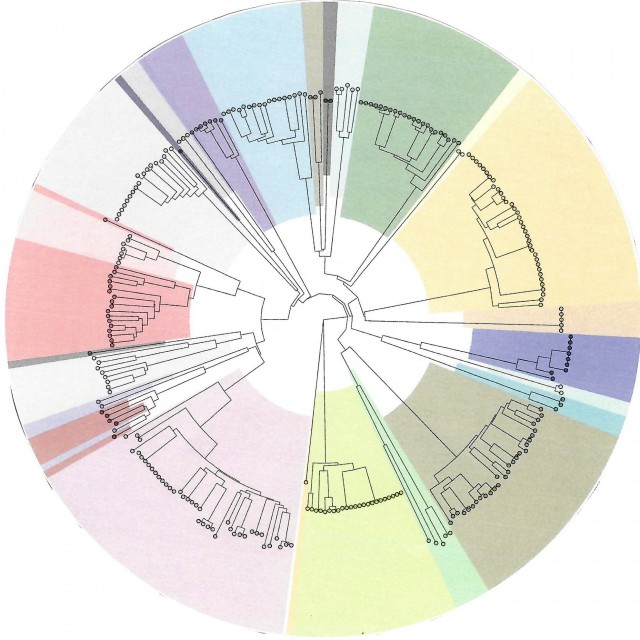
Duchossois Family Institute Microbiome Metabolomics (DFIMM) provides resources to measure, characterize, and identify the metabolites produced or modified by microbes to better understand how these molecules function in microbe-microbe and host-microbe interactions/communication and how these processes influence human health and disease. Visit the DFIMM website for information regarding services and pricing.
The Microbiome Metagenomics Facility (MMF) provides bacterial 16S ribosomal RNA gene and shotgun metagenomics sequence analyses on intestinal contents and other sample types containing complex microbial populations. Information on services and pricing can be found on the MMF website.
The Symbiotic Bacterial Strain Bank (SBSB) contains almost 2000 bacterial strains, isolated from healthy human fecal donors, that have been anaerobically cultured and characterized and are available to the research community. Information on ordering strains is provided on the SBSB website.
Supporting our facilities are our bioinformaticians who provide comprehensive computational data analyses, including genome assembly and annotation, using computation software programs to analyze microbial genome sequences and characterizing their metabolite utilization and secretion to better understand how commensal bacteria impact human health.